- Count the values of each class.
brain_df['mask'].value_counts()

- Randomly display an MRI image from the dataset.
image = cv2.imread(brain_df.image_path[1301]) plt.imshow(image)
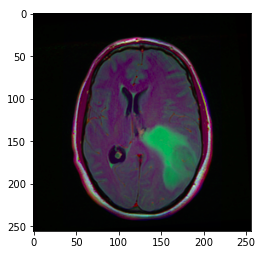
Image_path stores the path of the brain MRI so we can display the image using matplotlib.
Suggestion: the greenish part of the image above can be considered as the tumor.
- What's more, show the corresponding mask image.
image1 = cv2.imread(brain_df.mask_path[1301]) plt.imshow(image1)
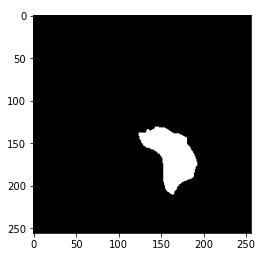
Now, you may have gotten the hint of what the mask really is. The mask is the image of the part of the brain affected by a tumor from the corresponding MRI image. Here, the mask is from the brain MRI shown above.
- Analyze the pixel values of the mask image.
cv2.imread(brain_df.mask_path[1301]).max()
Departure: 255
The maximum value of pixels in the mask image is 255, what the white color indicates.
cv2.imread(brain_df.mask_path[1301]).min()
Departure: 0
The minimum value of pixels in the mask image is 0, what the black color indicates.
- Visualization of MRI of the brain, the corresponding mask and MRI with the mask.
count = 0
fig, axs = plt.subplots(12, 3, figsize = (20, 50))
for i in range(len(brain_df)):
if brain_df['mask'][i] ==1 and count <5:
img = io.imread(brain_df.image_path[i])
axs[count][0].title.set_text('Brain MRI')
axs[count][0].imshow(img)
mask = io.imread(brain_df.mask_path[i])
axs[count][1].title.set_text('Mask')
axs[count][1].imshow(mask, cmap = 'gray')
img[mask == 255] = (255, 0, 0) #Red color
axs[count][2].title.set_text('MRI with Mask')
axs[count][2].imshow(img)
count+=1
fig.tight_layout()
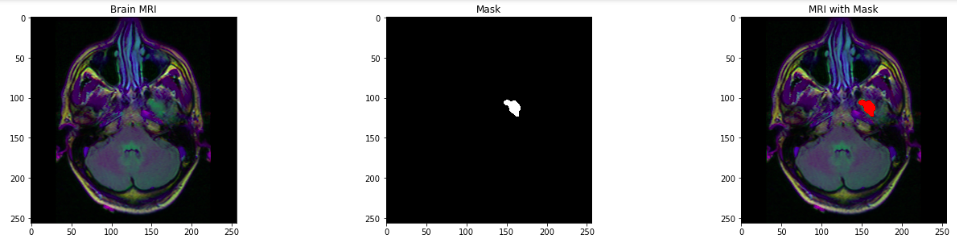
- Remove identification, as it is not necessary for processing.
# Drop the patient id column brain_df_train = brain_df.drop(columns = ['patient_id']) brain_df_train.shape
You will get the size of the data frame in the output: (3929, 3)
- Convert data in mask column from integer format to string format, since we will need the data in string format.
brain_df_train['mask'] = brain_df_train['mask'].apply(lambda x: str(x)) brain_df_train.info()
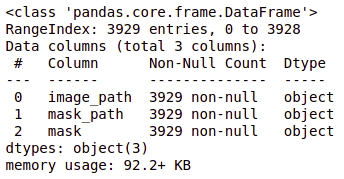
As you can see, now each feature has data type as object.
- Divide the data into test and train sets.
# split the data into train and test data from sklearn.model_selection import train_test_split train, test = train_test_split(brain_df_train, test_size = 0.15) |
- Increase more data with ImageDataGenerator. ImageDataGenerator generates batches of tensioner image data with real-time data augmentation.
Refer here for more information about ImageDataGenerator and parameters in detail.
We will create a train_generator and validation_generator from the train data and a test_generator from the test data.
# create an image generator from keras_preprocessing.image import ImageDataGenerator #Create a data generator which scales the data from 0 to 1 and makes validation split of 0.15 datagen = ImageDataGenerator(rescale=1./255., validation_split = 0.15) train_generator=datagen.flow_from_dataframe( dataframe=train, directory= './', x_col="image_path", y_col ="mask", subset="training", batch_size=16, shuffle=True, class_mode="categorical", target_size=(256,256)) valid_generator=datagen.flow_from_dataframe( dataframe=train, directory= './', x_col="image_path", y_col ="mask", subset="validation", batch_size=16, shuffle=True, class_mode="categorical", target_size=(256,256)) # Create a data generator for test images test_datagen=ImageDataGenerator(rescale=1./255.) test_generator=test_datagen.flow_from_dataframe( dataframe=test, directory= './', x_col="image_path", y_col ="mask", batch_size=16, shuffle=False, class_mode="categorical", target_size=(256,256))






